Generates comprehensive survival summary tables with survival probabilities at specified time points, median survival times, and optional group comparisons with statistical testing. Designed for creating survival summaries commonly used in clinical and epidemiological research publications.
Usage
survtable(
data,
outcome,
by = NULL,
times = NULL,
probs = 0.5,
stats = c("survival", "ci"),
type = "survival",
conf_level = 0.95,
conf_type = "log",
digits = 0,
time_digits = 1,
p_digits = 3,
percent = TRUE,
test = TRUE,
test_type = "logrank",
total = TRUE,
total_label = "Total",
time_unit = NULL,
time_label = NULL,
median_label = NULL,
labels = NULL,
by_label = NULL,
na_rm = TRUE,
number_format = NULL,
...
)Arguments
- data
Data frame or data.table containing the survival dataset. Automatically converted to a data.table for efficient processing.
- outcome
Character string or character vector specifying one or more survival outcomes using
Surv()syntax (e.g.,"Surv(os_months, os_status)"). When multiple outcomes are provided, results are stacked into a single table with outcome labels as row headers.- by
Character string specifying the column name of the stratifying variable for group comparisons (e.g., treatment arm, risk group). When
NULL(default), produces overall survival summaries only.- times
Numeric vector of time points at which to estimate survival probabilities. For example,
c(12, 24, 36)for 1-, 2-, and 3-year survival when time is measured in months. Default isNULL.- probs
Numeric vector of survival probabilities for which to estimate corresponding survival times (quantiles). Values must be between 0 and 1. For example,
c(0.5)returns median survival time,c(0.25, 0.5, 0.75)returns quartiles. Default is0.5(median only).- stats
Character vector specifying which statistics to display:
"survival"- Survival probability at specified times"ci"- Confidence interval for survival probability"n_risk"- Number at risk at each time point"n_event"- Cumulative number of events by each time point
Default is
c("survival", "ci").- type
Character string specifying the type of probability to report:
"survival"- Survival probability S(t) [default]"risk"- Cumulative incidence/risk 1 - S(t)"cumhaz"- Cumulative hazard -log(S(t))
- conf_level
Numeric confidence level for confidence intervals. Must be between 0 and 1. Default is 0.95 (95% confidence intervals).
- conf_type
Character string specifying the confidence interval type for survival estimates:
"log"- Log transformation (default, recommended)"log-log"- Log-log transformation"plain"- Linear/identity (can produce CIs outside [0, 1])"logit"- Logit transformation"arcsin"- Arcsin square root transformation
- digits
Integer specifying the number of decimal places for survival probabilities (as percentages). Default is 0 (whole percentages).
- time_digits
Integer specifying the number of decimal places for survival time estimates (median, quantiles). Default is 1.
- p_digits
Integer specifying the number of decimal places for p-values. Values smaller than
10^(-p_digits)are displayed as"< 0.001"(forp_digits = 3),"< 0.0001"(forp_digits = 4), etc. Default is 3.- percent
Logical. If
TRUE(default), displays survival probabilities as percentages (e.g.,"85%"). IfFALSE, displays as proportions (e.g.,"0.85").- test
Logical. If
TRUE(default), performs a survival curve comparison test and adds a p-value column. Requiresbyto be specified.- test_type
Character string specifying the statistical test for comparing survival curves:
"logrank"- Log-rank test (default)"wilcoxon"- Wilcoxon (Breslow) test"tarone"- Tarone-Ware test"petopeto"- Peto-Peto test
- total
Logical or character string controlling the total/overall column:
TRUEor"first"- Include total column first [default]"last"- Include total column last (before p-value)FALSE- Exclude total column
- total_label
Character string for the total/overall row label. Default is
"Total".- time_unit
Character string specifying the time unit for display in column headers and labels (e.g.,
"months","days","years"). When specified, time column headers become "{time} {time_unit}" (e.g., "12 months"). Default isNULL(no unit shown).- time_label
Character string template for time column headers when
timesis specified. Use"\{time\}"as placeholder for the time value and"\{unit\}"for the time unit. Default is"\{time\} \{unit\}"whentime_unitis specified, otherwise just"\{time\}".- median_label
Character string for the median survival row label. Default is
NULL, which auto-constructs fromconf_level(e.g.,"Median (95% CI)"forconf_level = 0.95).- labels
Named character vector or list providing custom display labels. For stratified analyses, names should match levels of the
byvariable. For multiple outcomes, names should match theSurv()expressions. Default isNULL.- by_label
Character string providing a custom label for the stratifying variable (used in output attributes and headers). Default is
NULL(uses variable name).- na_rm
Logical. If
TRUE(default), observations with missing values in time, status, or the stratifying variable are excluded.- number_format
Character string or two-element character vector controlling thousand and decimal separators in formatted output. Named presets:
"us"- Comma thousands, period decimal:1,234.56[default]"eu"- Period thousands, comma decimal:1.234,56"space"- Thin-space thousands, period decimal:1 234.56(SI/ISO 31-0)"none"- No thousands separator:1234.56
Or provide a custom two-element vector
c(big.mark, decimal.mark), e.g.,c("'", ".")for Swiss-style:1'234.56.When
NULL(default), usesgetOption("summata.number_format", "us"). Set the global option once per session to avoid passing this argument repeatedly:options(summata.number_format = "eu")- ...
Additional arguments passed to
survfit.
Value
A data.table with S3 class "survtable" containing formatted
survival statistics. The table structure depends on parameters:
When times is specified (survival at time points):
- Variable/Group
Row identifier – stratifying variable levels
- Time columns
Survival statistics at each requested time point
- p-value
Test p-value (if
test = TRUEandbyspecified)
When only probs is specified (survival quantiles):
- Variable/Group
Row identifier – stratifying variable levels
- Quantile columns
Time to reach each survival probability
- p-value
Test p-value (if
test = TRUEandbyspecified)
All numeric output (probabilities, times, counts, p-values)
respects the number_format setting for locale-appropriate
formatting.
The returned object includes the following attributes:
- raw_data
Data.table with unformatted numeric values
- survfit_objects
List of survfit objects for each stratum
- by_variable
The stratifying variable name
- times
The time points requested
- probs
The probability quantiles requested
- test_result
Full test result object (if test performed)
Details
Survival Probability Estimation:
Survival probabilities are estimated using the Kaplan-Meier method via
survfit. At each specified time point, the function
reports the estimated probability of surviving beyond that time.
Confidence Intervals:
The default "log" transformation for confidence intervals is
recommended as it ensures intervals remain within [0, 1] and has good
statistical properties. The "log-log" transformation is also
commonly used and may perform better in the tails.
Statistical Testing:
The log-rank test (default) tests the null hypothesis that survival curves are identical across groups. Alternative tests weight different parts of the survival curve:
Log-rank: Equal weights (best for proportional hazards)
Wilcoxon: Weights by number at risk (sensitive to early differences)
Tarone-Ware: Weights by square root of number at risk
Peto-Peto: Modified Wilcoxon weights
Formatting:
All numeric output respects the number_format parameter.
Separators within confidence intervals adapt automatically to avoid
ambiguity:
Survival probabilities:
"85% (80%-89%)"(US) or"85% (80%-89%)"(EU, en-dash separator)Median survival:
"24.5 (21.2-28.9)"(US) or"24,5 (21,2-28,9)"(EU)Counts \(\ge\) 1000:
"1,234"(US) or"1.234"(EU)p-values:
"< 0.001"(US) or"< 0,001"(EU)
See also
desctable for baseline characteristics tables,
fit for regression analysis,
table2pdf for PDF export,
table2docx for Word export,
survfit for underlying survival estimation,
survdiff for survival curve comparison tests
Other descriptive functions:
desctable(),
print.survtable()
Examples
# Load example data
data(clintrial)
# Example 1: Survival at specific time points by treatment
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24, 36),
time_unit = "months"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24, 36 months
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 months 24 months 36 months Median (95% CI) p-value
#> <char> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (43-49%) 36% (33-40%) 19.4 (16.2-23.4) < 0.001
#> 2: Control 55% (49-63%) 37% (31-44%) 26% (20-33%) 14.7 (10.5-19.2)
#> 3: Drug A 68% (63-74%) 56% (51-62%) 48% (42-54%) 33.6 (24.5-42.2)
#> 4: Drug B 54% (49-59%) 43% (38-48%) 33% (28-38%) 14.7 (11.0-21.8)
# \donttest{
# Example 2: Median survival only
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = NULL,
probs = 0.5
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable Median (95% CI) p-value
#> <char> <char> <char>
#> 1: Total 19.4 (16.2-23.4) < 0.001
#> 2: Control 14.7 (10.5-19.2)
#> 3: Drug A 33.6 (24.5-42.2)
#> 4: Drug B 14.7 (11.0-21.8)
# Example 3: Multiple quantiles (quartiles)
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "stage",
times = NULL,
probs = c(0.25, 0.5, 0.75)
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: stage
#> Quantiles: 25%, 50%, 75%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 75% Survival Time (95% CI) Median (95% CI) 25% Survival Time (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 5.1 (4.3-6.1) 19.5 (16.1-23.5) NR < 0.001
#> 2: I 9.0 (7.4-12.5) 33.7 (22.4-45.2) NR
#> 3: II 7.7 (6.1-12.3) 32.2 (24.8-42.0) NR
#> 4: III 3.9 (3.0-5.2) 13.7 (10.4-22.3) 44.7 (35.8-NR)
#> 5: IV 2.0 (1.4-2.8) 5.9 (4.3-8.2) 15.6 (10.8-26.7)
# Example 4: Both time points and median
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
probs = 0.5,
time_unit = "months"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24 months
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 months 24 months Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (43-49%) 19.4 (16.2-23.4) < 0.001
#> 2: Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2)
#> 3: Drug A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 4: Drug B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 5: Cumulative incidence (1 - survival)
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
type = "risk"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Cumulative incidence
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 41% (38-44%) 54% (51-57%) 19.4 (16.2-23.4) < 0.001
#> 2: Control 45% (37-51%) 63% (56-69%) 14.7 (10.5-19.2)
#> 3: Drug A 32% (26-37%) 44% (38-49%) 33.6 (24.5-42.2)
#> 4: Drug B 46% (41-51%) 57% (52-62%) 14.7 (11.0-21.8)
# Example 6: Include number at risk
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
stats = c("survival", "ci", "n_risk")
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) [n = 488] 46% (43-49%) [n = 369] 19.4 (16.2-23.4) < 0.001
#> 2: Control 55% (49-63%) [n = 106] 37% (31-44%) [n = 70] 14.7 (10.5-19.2)
#> 3: Drug A 68% (63-74%) [n = 192] 56% (51-62%) [n = 154] 33.6 (24.5-42.2)
#> 4: Drug B 54% (49-59%) [n = 190] 43% (38-48%) [n = 145] 14.7 (11.0-21.8)
# Example 7: Overall survival without stratification
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
times = c(12, 24, 36, 48)
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Time points: 12, 24, 36, 48
#> Quantiles: 50%
#> Statistic: Survival probability
#>
#> Variable 12 24 36 48 Median (95% CI)
#> <char> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (43-49%) 36% (33-40%) 30% (27-33%) 19.4 (16.2-23.4)
# Example 8: Without total row
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
total = FALSE
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2) < 0.001
#> 2: Drug A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 3: Drug B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 9: Custom labels
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
labels = c("Drug A" = "Treatment A", "Drug B" = "Treatment B"),
time_unit = "months"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24 months
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 months 24 months Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (43-49%) 19.4 (16.2-23.4) < 0.001
#> 2: Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2)
#> 3: Treatment A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 4: Treatment B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 10: Different confidence interval type
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
conf_type = "log-log"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (42-49%) 19.4 (16.1-22.9) < 0.001
#> 2: Control 55% (48-62%) 37% (30-44%) 14.7 (10.2-19.0)
#> 3: Drug A 68% (63-73%) 56% (50-62%) 33.6 (24.4-40.9)
#> 4: Drug B 54% (48-59%) 43% (38-48%) 14.7 (10.9-21.7)
# Example 11: Wilcoxon test instead of log-rank
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
test_type = "wilcoxon"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Wilcoxon (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56-62%) 46% (43-49%) 19.4 (16.2-23.4) < 0.001
#> 2: Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2)
#> 3: Drug A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 4: Drug B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 12: Access raw data for custom analysis
result <- survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24)
)
raw <- attr(result, "raw_data")
print(raw)
#> Variable 12_estimate 12_lower 12_upper 12_n_risk 12_n_event 24_estimate 24_lower 24_upper 24_n_risk 24_n_event Median (95% CI)_estimate Median (95% CI)_lower Median (95% CI)_upper
#> <char> <num> <num> <num> <num> <num> <num> <num> <num> <num> <num> <num> <num> <num>
#> 1: Total 0.5897650 0.5573977 0.6240118 488 345 0.4589077 0.4262187 0.4941037 369 107 19.4 16.2 23.4
#> 2: Control 0.5518225 0.4860549 0.6264890 106 87 0.3686484 0.3063818 0.4435696 70 35 14.7 10.5 19.2
#> 3: Drug A 0.6829195 0.6310704 0.7390285 192 91 0.5604338 0.5054384 0.6214131 154 34 33.6 24.5 42.2
#> 4: Drug B 0.5361330 0.4869622 0.5902688 190 167 0.4272844 0.3788226 0.4819458 145 38 14.7 11.0 21.8
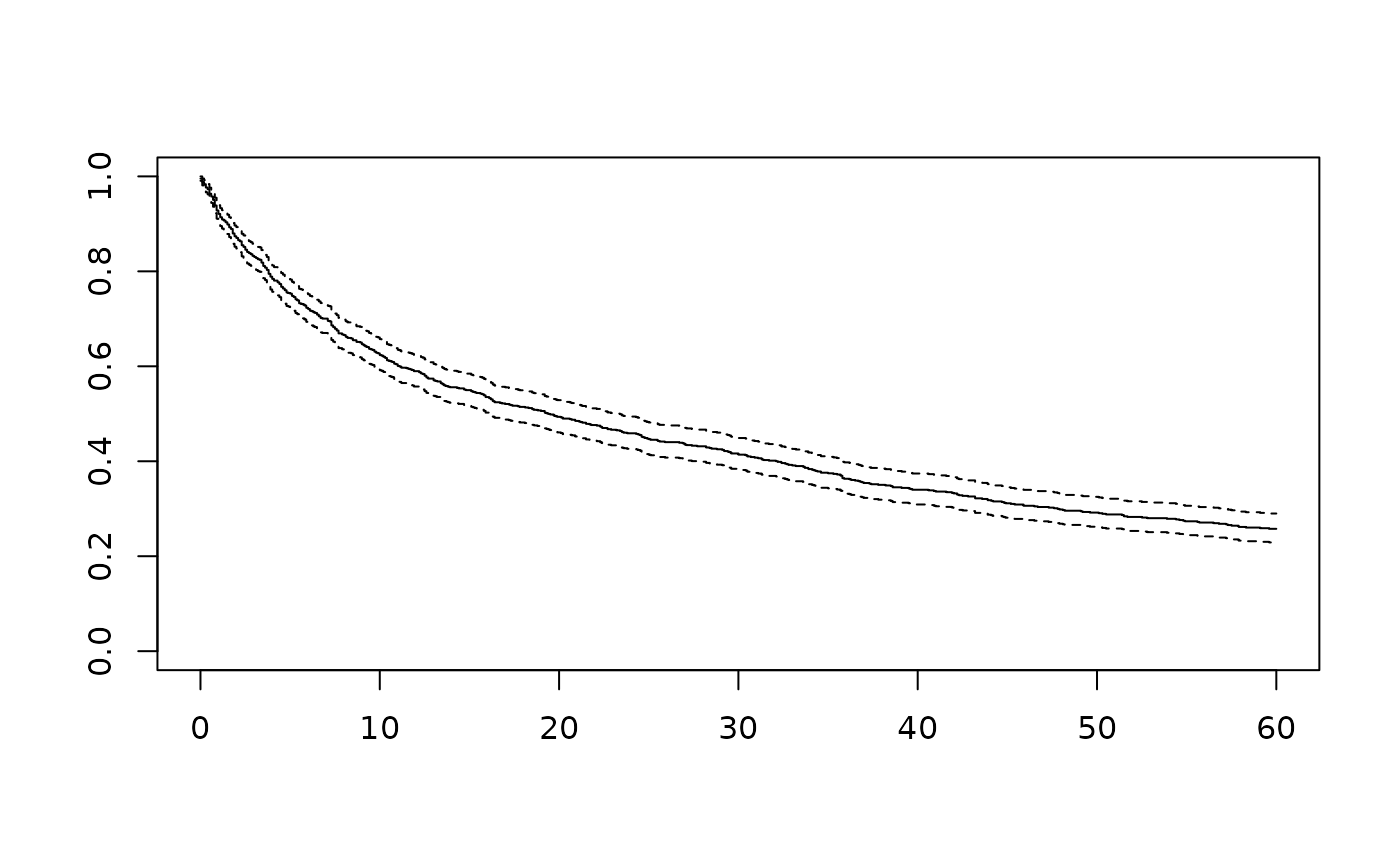
# Example 13: Access survfit objects for plotting
fits <- attr(result, "survfit_objects")
plot(fits$overall) # Plot overall survival curve
 # Example 14: Multiple survival outcomes stacked
survtable(
data = clintrial,
outcome = c("Surv(pfs_months, pfs_status)", "Surv(os_months, os_status)"),
by = "treatment",
times = c(12, 24),
probs = 0.5,
time_unit = "months",
total = FALSE,
labels = c(
"Surv(pfs_months, pfs_status)" = "Progression-Free Survival",
"Surv(os_months, os_status)" = "Overall Survival"
)
)
#>
#> Survival Summary Table
#> Outcomes: 2
#> Stratified by: treatment
#> Time points: 12, 24 months
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank
#>
#> Variable Group 12 months 24 months Median (95% CI) p-value
#> <char> <char> <char> <char> <char> <char>
#> 1: Progression-Free Survival Control 29% (23-36%) 14% (9-20%) 4.7 (3.6-6.9) < 0.001
#> 2: Drug A 44% (39-51%) 30% (25-36%) 9.0 (7.5-12.4)
#> 3: Drug B 30% (26-36%) 18% (14-22%) 4.7 (3.7-6.2)
#> 4: Overall Survival Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2) < 0.001
#> 5: Drug A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 6: Drug B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 15: European number formatting
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
number_format = "eu"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56–62%) 46% (43–49%) 19,4 (16,2–23,4) < 0,001
#> 2: Control 55% (49–63%) 37% (31–44%) 14,7 (10,5–19,2)
#> 3: Drug A 68% (63–74%) 56% (51–62%) 33,6 (24,5–42,2)
#> 4: Drug B 54% (49–59%) 43% (38–48%) 14,7 (11,0–21,8)
# }
# Example 14: Multiple survival outcomes stacked
survtable(
data = clintrial,
outcome = c("Surv(pfs_months, pfs_status)", "Surv(os_months, os_status)"),
by = "treatment",
times = c(12, 24),
probs = 0.5,
time_unit = "months",
total = FALSE,
labels = c(
"Surv(pfs_months, pfs_status)" = "Progression-Free Survival",
"Surv(os_months, os_status)" = "Overall Survival"
)
)
#>
#> Survival Summary Table
#> Outcomes: 2
#> Stratified by: treatment
#> Time points: 12, 24 months
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank
#>
#> Variable Group 12 months 24 months Median (95% CI) p-value
#> <char> <char> <char> <char> <char> <char>
#> 1: Progression-Free Survival Control 29% (23-36%) 14% (9-20%) 4.7 (3.6-6.9) < 0.001
#> 2: Drug A 44% (39-51%) 30% (25-36%) 9.0 (7.5-12.4)
#> 3: Drug B 30% (26-36%) 18% (14-22%) 4.7 (3.7-6.2)
#> 4: Overall Survival Control 55% (49-63%) 37% (31-44%) 14.7 (10.5-19.2) < 0.001
#> 5: Drug A 68% (63-74%) 56% (51-62%) 33.6 (24.5-42.2)
#> 6: Drug B 54% (49-59%) 43% (38-48%) 14.7 (11.0-21.8)
# Example 15: European number formatting
survtable(
data = clintrial,
outcome = "Surv(os_months, os_status)",
by = "treatment",
times = c(12, 24),
number_format = "eu"
)
#>
#> Survival Summary Table
#> Outcome: Surv(os_months, os_status)
#> Stratified by: treatment
#> Time points: 12, 24
#> Quantiles: 50%
#> Statistic: Survival probability
#> Test: Log-rank (p = < 0.001)
#>
#> Variable 12 24 Median (95% CI) p-value
#> <char> <char> <char> <char> <char>
#> 1: Total 59% (56–62%) 46% (43–49%) 19,4 (16,2–23,4) < 0,001
#> 2: Control 55% (49–63%) 37% (31–44%) 14,7 (10,5–19,2)
#> 3: Drug A 68% (63–74%) 56% (51–62%) 33,6 (24,5–42,2)
#> 4: Drug B 54% (49–59%) 43% (38–48%) 14,7 (11,0–21,8)
# }